Bonus topic: Add Adsorbate#
Introduction#
In many calculations we use an adsorbate-substrate complex. In the old days, we manually create the location of the adsorbate on top of the substrate (via Excel or by-hand). Now we can use ASE to do this for us. The ASE one is not simple to use, so I created my own wrapper of it.
Installation#
pip install git+https://github.com/kimrojas/MatSciToolKit.git
Usage#
Simple case#
I have two trajectory/structure file: nh3.traj (adsorbate) and hb.traj (substrate). Data files available here.
from ase.io import read, write
from matscitoolkit.builder import add_adsorbate
from ase.visualize.plot import plot_atoms
from pathlib import Path
import matplotlib.pyplot as plt
cwd = Path.cwd()
bookdir = list(cwd.parents)[2]
filedir = bookdir / "files/qe_tutorial/bonus_add_adsorbate"
# Load the two structures
adsorbate = read(filedir / "nh3.traj")
substrate = read(filedir / "hb.traj")
# Specify the height
height = 3.
# Add the adsorbate
atm_complex = add_adsorbate(adsorbate, substrate, height,
substrate_reference=[11, 20], # Mid point between substrate atom 11 and 20
adsorbate_origin="COG", # Center of geometry of the adsorbate (other opt: `COM` or specific atom index)
adsorbate_rotation=None, # Specify the rotation if needed
)
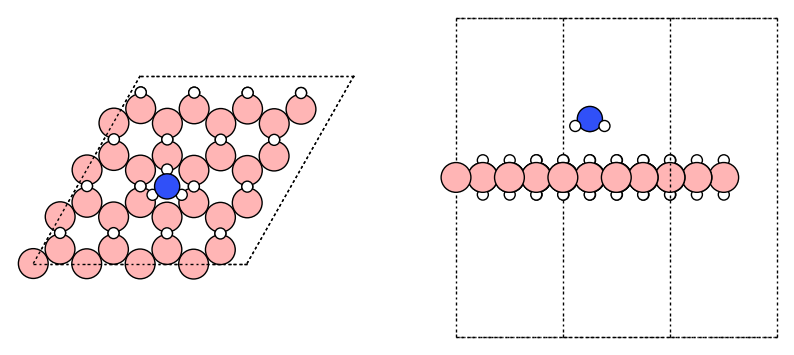
fig, ax = plt.subplots(1, 2, figsize=(10, 5))
plot_atoms(atm_complex, ax[0]);
plot_atoms(atm_complex, ax[1], rotation="-90x");
for iax in ax:
iax.axis("off")
Building structure at height: 3.0
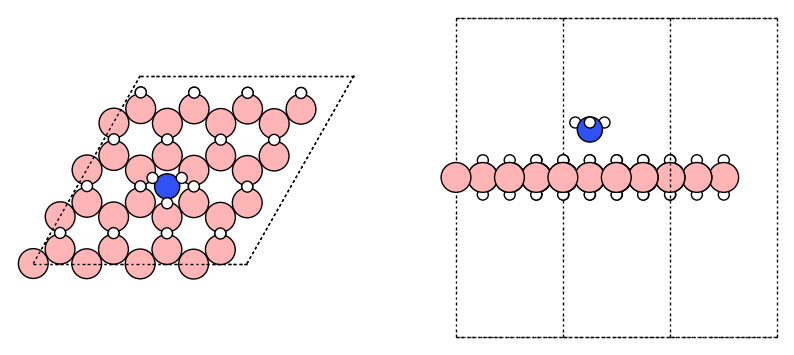
# Add the adsorbate with flip rotation
atm_complex = add_adsorbate(adsorbate, substrate, height,
substrate_reference=[11, 20], # Mid point between substrate atom 11 and 20
adsorbate_origin="COG", # Center of geometry of the adsorbate (other opt: `COM` or specific atom index)
adsorbate_rotation=[(180, 'x')], # Specify the rotation if needed
)
fig, ax = plt.subplots(1, 2, figsize=(10, 5))
plot_atoms(atm_complex, ax[0]);
plot_atoms(atm_complex, ax[1], rotation="-90x");
for iax in ax:
iax.axis("off")
Building structure at height: 3.0
More information#
For more information you can visit the documentation